Mitonuclear adapatation in eastern yellow robins
|
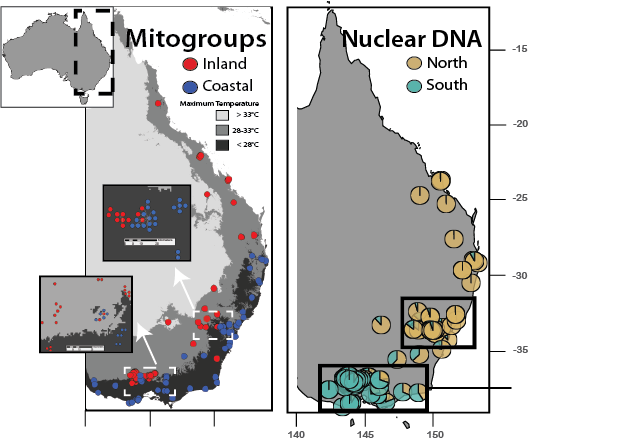
Eastern yellow robin represent one of the most striking examples of mitonuclear discordance (Pavlova et al. 2013):
a) Two deeply divergent (~6.6%) mitochondrial groups (i.e. mitogroups): inland and coastal. b) Distribution of mitogroups correlates with climatic variation after correcting for geography c) Mitogroups overlap but mtDNA mixing is minimal, despite the absence of physical barriers d) Nuclear genetic structure is minimal and perpendicular (i.e. discordant) to that of mtDNA e) Sex-biased dispersal, vicariance or incomplete lineage sorting cannot explain the divergence, so it is likely product of female-associated selection |
I'm interested in resolving how mitochondrial genes of yellow robins can be so divergent while undergoing gene flow. Using complete mitochondrial genomes we found molecular evidence of positive selection acting on few amino acid sites of the mitochondrial genome against a background of strong purifying selection (Morales et al. 2015).
Now I endeavour to look for nuclear genes that might follow the same pattern. Using a reduce genome representation scan via diversity arrays (i.e. DArTs) I'm looking for SNPs exceptionally different between mitogroups and also those that correlate with key environmental variables. Preliminar results will be presented in the coming European Society of Evolutionary Biology Conference (check out the poster here)
Members of the highly divergent groups besides undergoing extensive gene flow do not have any obvious morphological differentiation. However, the extreme observed divergence is maintained. I'm going to employ morphological, plumage colour variation, song recordings and hundreds of sequence based markers to gain insight into potential pre- and post-zygotic barriers between mitogroups
Now I endeavour to look for nuclear genes that might follow the same pattern. Using a reduce genome representation scan via diversity arrays (i.e. DArTs) I'm looking for SNPs exceptionally different between mitogroups and also those that correlate with key environmental variables. Preliminar results will be presented in the coming European Society of Evolutionary Biology Conference (check out the poster here)
Members of the highly divergent groups besides undergoing extensive gene flow do not have any obvious morphological differentiation. However, the extreme observed divergence is maintained. I'm going to employ morphological, plumage colour variation, song recordings and hundreds of sequence based markers to gain insight into potential pre- and post-zygotic barriers between mitogroups
Plumage colour variation
To complicate things further, yellow robins also show a pattern of plumage colour variation that is spatially concordant with the nuclear differentiation but not with the mitochondrial divergence. Various hypothesis have been formulated to explain this pattern, including those that suggest that is maintained by natural selection imposed by visual environments (open vs. closed vegetation). I plan to tackle this question comparing the colour variation with environmental and neutral genetic data to estimate which driver is most likely maintaining the colour variation.

Plumage-colour variation and examples of the most common habitat types in each region. Data is from Keast 1958: Rec Aust Mus; and Ford 1979: Emu. Map was created by Nevil Amos. Bird photos by Chris Tzaros. North habitat corresponds to Wooroonooran National Park (Photo by
Aleksandra Srsa). South habitat corresponds to Holey Plains State Park (Photo by Peter Whitehead).
Aleksandra Srsa). South habitat corresponds to Holey Plains State Park (Photo by Peter Whitehead).